Biomedical
A New Antibiotic Target for Gram-Positive Bacteria
The technology
Clinical treatment of S. aureus infections has been compromised by the rapid resistance to multiple antibiotics. Methicillin-resistant S. aureus (MRSA) is one such strain which can cause severe complications and death. Identification of novel molecular targets is seen as a major obstacle to the development of new antibiotics for MRSA infections. There is a clinical need for a treatment of infection that targets S. aureus which have acquired resistance to other medications.
Technology summary
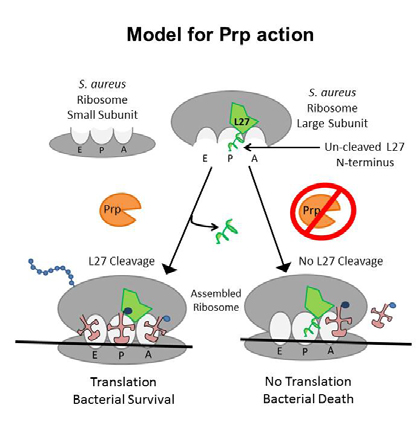
VCU researchers have identified and performed the initial characterization of an essential, highly conserved protease unique to S. aureus and other Gram-positive pathogens such as Bacillus, Clostridium, and Streptococcus. This protease, Prp, performs a novel site-specific cleavage of ribosomal protein L27 that is essential for bacterial survival. Lack of cleavage by this protease either prevents proper ribosome assembly or blocks peptidyl transferase activity. An assay has been developed that is suitable for high throughput screening of compounds that inhibit the activity of this enzyme. Thus, this protease could be a prime target for novel antibiotics specific to Staphylococcus and other Gram-positive bacteria.
Technology status
Edman degradation has been performed to confirm cleavage by the protease and the crystal structure of the protease has been reported. Molecular modeling for the purposes of drug design has been performed.
Michael S. Spilman et al. “Assembly of bacteriophage 80α capsids in a Staphylococcus aureus expression system”. Virology, 2012, 434(2), 242-250.
Erin A. Wall et al. “Specific N-terminal cleavage of ribosomal protein L27 in Staphylococcus aureus and related bacteria”. Molecular Biology, 2014, 95(2), 258-269
